nf-core/proteinfold
Protein 3D structure prediction pipeline
Introduction
This document describes the user-facing output produced by the pipeline.
Pipeline overview
The pipeline is built using Nextflow and predicts protein structures using the following methods:
See main README.md for a condensed overview of the steps in the pipeline, and the bioinformatics tools used at each step.
The directories listed below will be created in the output directory after the pipeline has finished. All paths are relative to the top-level results directory.
Exact subdirectories depend on the selected mode(s). In a multi-mode run (for example alphafold2,boltz,rosettafold_all_atom) you will typically see top-level directories such as alphafold2/, boltz/, rosettafold_all_atom/, multiqc/, reports/, compare/, and pipeline_info/.
Prediction outputs (all modes)
User-facing outputs are largely consistent across modes.
Common output patterns
<MODE>/top_ranked_structures/<SEQUENCE NAME>.pdb<MODE>/<SEQUENCE NAME>/<SEQUENCE NAME>_plddt.tsv<MODE>/<SEQUENCE NAME>/paes/<SEQUENCE NAME>_<RANK>_pae.tsv(when available)<MODE>/<SEQUENCE NAME>/<SEQUENCE NAME>_<MODE>_msa.tsv(mode-specific MSA summary)<MODE>/<SEQUENCE NAME>/<SEQUENCE NAME>_{ptm,iptm}.tsvand chainwise summaries (where applicable)
pLDDT ({meta.id}_plddt.tsv)
Confidence values per residue, rounded to 2 decimal places. Each ranked result gets its own column (for all-atom modules, atomic token confidences are processed to a naive mean value across the residue).
Positions rank_0 rank_1 rank_2 rank_3 rank_4
0 83.58 85.27 88.41 86.22 84.91
1 97.99 97.81 97.39 97.49 97.32
2 98.22 98.42 98.16 97.88 97.81
3 98.06 98.15 97.94 97.56 97.4
4 98.67 98.56 98.3 98.38 98.29
5 98.81 98.77 98.62 98.61 98.54
6 98.79 98.74 98.57 98.59 98.52
...MSA ({meta.id}_{meta.mode}_msa.tsv)
The amino acid characters are converted to integers 0-19, unknown as 20, integer 21 represents the gap character.
19 5 5 4 10 16 15 3 8 15 13 16 12 9 17 16 9 4 8 11 0 7 7 8 11 0 19 8 8 5 3
19 5 5 4 10 16 15 3 8 15 13 16 12 9 17 16 9 4 8 11 0 7 7 8 11 0 6 8 8 5 13
19 5 5 4 10 5 15 13 14 0 14 16 12 9 17 16 9 4 14 11 0 7 5 8 15 4 5 8 3 5 21
19 5 5 4 10 16 15 3 8 15 13 16 12 9 17 16 9 4 8 11 0 7 7 8 11 0 19 8 8 5 21
19 5 5 4 10 16 15 3 8 15 13 16 12 9 17 16 9 4 8 11 0 7 7 8 11 0 19 8 8 5 13
19 5 5 4 10 16 15 3 8 15 13 16 12 9 7 16 9 4 8 11 0 7 7 8 11 0 6 8 8 5 13This allows easy sequence indentity calculation when processing as a numpy array.
(i)pTM ({meta.id}_[i]ptm.tsv)
(i)pTM scores, rounded to 3 decimal places, listed by the rank number (currently unsorted - to reflect models and seeds where appropriate).
17 0.552
22 0.529
21 0.532
20 0.541
23 0.523
3 0.606
2 0.610
4 0.606
1 0.616
0 0.617
12 0.580
9 0.588
13 0.580
11 0.583
14 0.570
15 0.565
24 0.517
16 0.560
18 0.550
19 0.550
10 0.588
5 0.600
6 0.597
7 0.596
8 0.595chain-wise (i)pTM ({meta.id}_chainwise_[i]ptm.tsv)
(Asymmetrical) ipTM scores, rounded to 4 decimal places, with chain pair lettering as the row (X:Y), and the rank number as the column. A pTM value is a chain’s own predicted Template Modelling score so lettering will be X:X.
0 1 2
A:B 0.2880 0.2750 0.2900
B:A 0.2904 0.2801 0.2915PAE ({meta.id}_{rank_number}_pae.tsv)
Predicted alignment error of residues j aligned by residue i, rounded to 4 decimal places.
The row number gives you the index of residue i and the column value within the row gives the index of residue j for the 2D PAE matrix.
Each model prediction generates a separate file containing the rank number. The _0_pae.tsv file corresponds to the top ranked model, other ranked results are stored within the paes/ folder.
0.2500 1.5710 3.9037 6.2177 8.4471 11.4583 12.9679 15.1237 18.0263 18.3868 18.9381 20.5747 19.3314 20.1825 21.6145 23.2190
2.2177 0.2500 1.5559 4.0327 6.3151 7.6372 10.1969 11.3626 14.9366 16.1303 17.9119 19.1877 21.2715 20.9531 20.1760 19.4087
3.4270 1.5284 0.2500 2.1333 3.5351 5.1049 6.6521 8.2317 12.1379 13.7185 14.9523 16.6154 19.6988 21.7614 18.6592 17.9619
6.1051 5.4206 2.5987 0.2500 2.0724 5.1454 6.7492 9.5538 9.6285 12.3868 13.8527 16.3586 17.2605 20.6381 19.9987 19.3295
7.3512 6.4947 5.5435 2.6740 0.2500 1.7561 4.9041 6.3923 8.9735 8.9272 12.3419 14.6005 15.9820 17.6358 20.5190 19.1028
7.4734 7.0899 5.8128 5.7512 2.0439 0.2500 1.8352 5.1064 6.4225 9.2098 10.5136 12.9404 14.3152 16.8122 18.6336 17.7382Example report plots
The report exports include key visualisations such as sequence coverage, predicted Local Distance Difference Test (pLDDT), and Predicted Aligned Error (PAE).
Sequence coverage
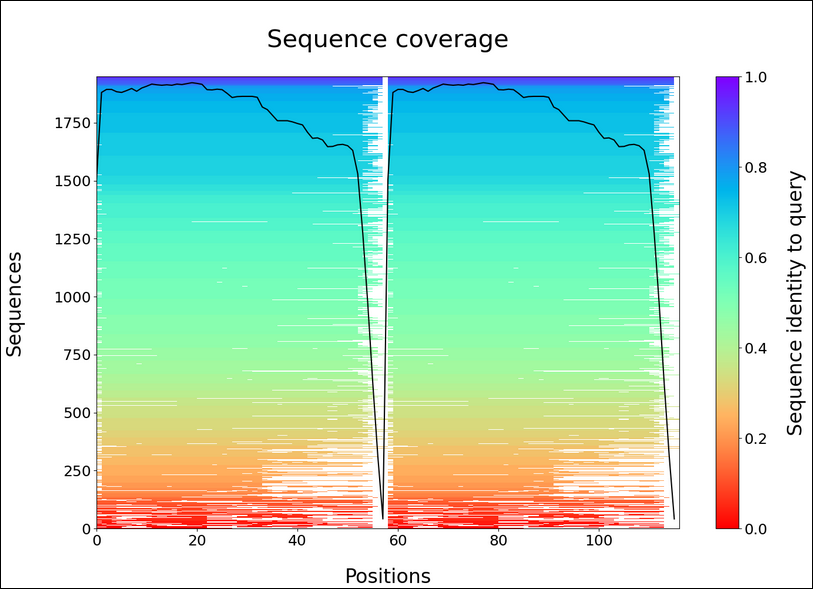
predicted Local Distance Difference Test (pLDDT)
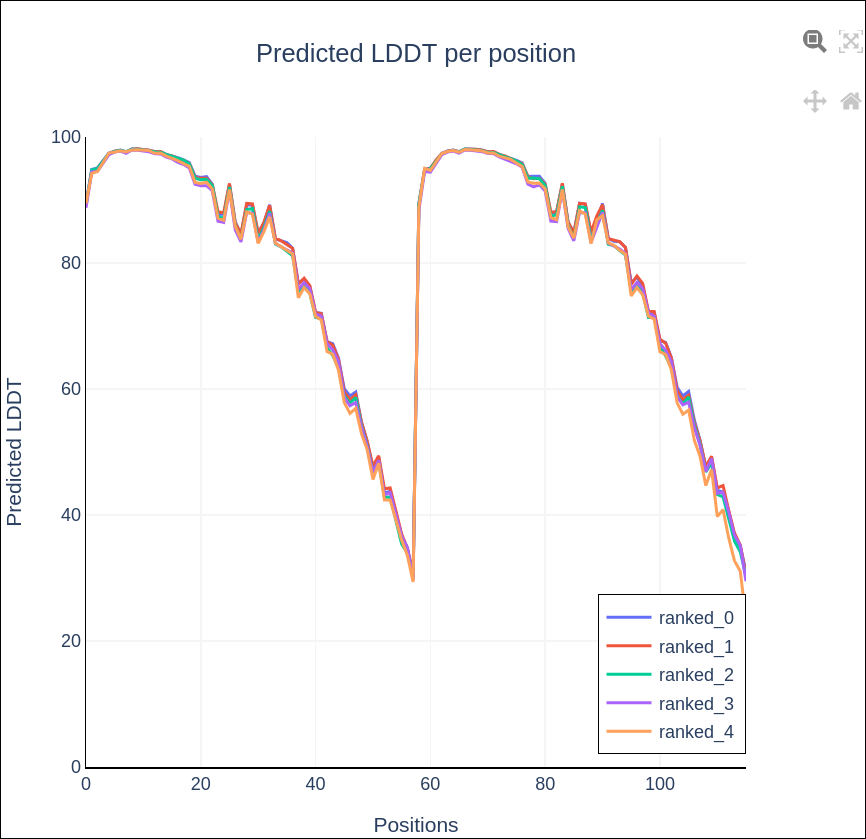
Predicted Aligned Error (PAE)
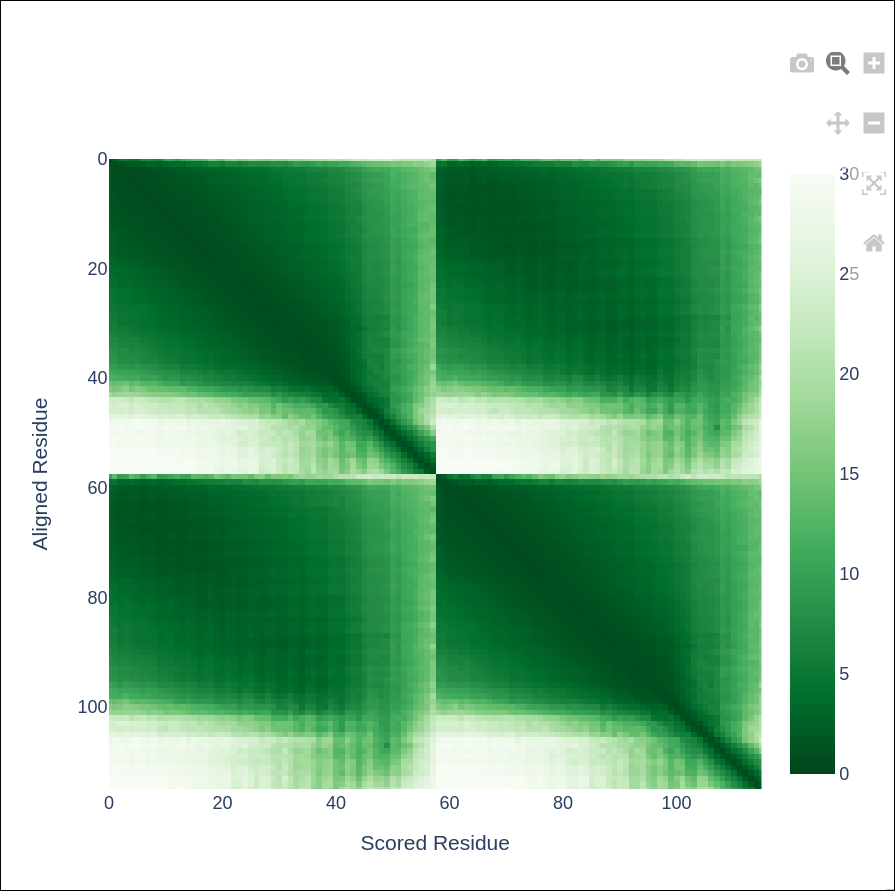
Per-mode reports and comparisons
Output files
reports/<SEQUENCE NAME>_<MODE>_report.html(single-mode report per sequence/mode)
compare/<SEQUENCE NAME>_comparison_report.html(present when running multiple modes)
Foldseek structural similarity search
If Foldseek is enabled (--skip_foldseek false), results are written to:
Output files
foldseek_easysearch/<SEQUENCE NAME>_<MODE>_foldseek.html(default output format)<SEQUENCE NAME>.m8(tabular output when--foldseek_easysearch_argdoes not include--format-mode 3)
Foldseek runs on top-ranked structures from each selected mode and sequence. By default, the pipeline uses --format-mode 3 and publishes HTML reports.
MultiQC report
Output files
multiqc*_multiqc_report.html: Standalone HTML report(s) that can be viewed in your web browser.*_multiqc_report_data/: Parsed report data for each corresponding MultiQC report.
MultiQC is a visualisation tool that generates HTML report(s) summarising samples in your project. Most QC results are visualised in the report and further statistics are available within each corresponding *_multiqc_report_data/ directory.
Results generated by MultiQC collate QC metrics from the selected structure-prediction mode(s), and the software versions for traceability. For more information about how to use MultiQC reports, see http://multiqc.info.
Pipeline information
Output files
pipeline_info/- Reports generated by Nextflow:
execution_report.html,execution_timeline.html,execution_trace.txtandpipeline_dag.dot/pipeline_dag.svg. - Reports generated by the pipeline:
pipeline_report.html,pipeline_report.txtandsoftware_versions.yml. Thepipeline_report*files will only be present if the--email/--email_on_failparameter’s are used when running the pipeline. - Reformatted samplesheet files used as input to the pipeline:
samplesheet.valid.csv. - Parameters used by the pipeline run:
params.json.
- Reports generated by Nextflow:
Nextflow provides excellent functionality for generating various reports relevant to the running and execution of the pipeline. This will allow you to troubleshoot errors with the running of the pipeline, and also provide you with other information such as launch commands, run times and resource usage.
Additional intermediate outputs
Depending on the selected mode(s) and options, additional top-level directories may be present, for example:
fasta2yaml/(for YAML conversion inputs/outputs)mmseqs/results/(for MMseqs2 outputs such as.a3mfiles)split/output_msa/(for split-MSA intermediate CSV outputs)
--save_intermediates
If --save_intermediates is enabled, extra raw intermediate files are published in mode-specific raw/ directories.
Examples include:
alphafold2/<MODE>/<SEQUENCE NAME>/raw/colabfold/<SEQUENCE NAME>/raw/boltz/<SEQUENCE NAME>/boltz_results_*/rosettafold_all_atom/<SEQUENCE NAME>/raw/alphafold3/<SEQUENCE NAME>/raw/helixfold3/<SEQUENCE NAME>/raw/rosettafold2na/<SEQUENCE NAME>/raw/
These raw outputs are intended for advanced debugging, reproducibility and method-specific downstream analyses. For detailed, canonical tool-specific native output specifications, see: