nf-core/bactmap
A mapping-based pipeline for creating a phylogeny from bacterial whole genome sequences
Introduction
nf-core/bactmap is a bioinformatics best-practice analysis pipeline for mapping short reads from bacterial WGS to a reference sequence, creating filtered VCF files, making pseudogenomes based on high quality positions in the VCF files and optionally creating a phylogeny from an alignment of the pseudogenomes.
The pipeline is built using Nextflow, a workflow tool to run tasks across multiple compute infrastructures in a very portable manner. It uses Docker/Singularity containers making installation trivial and results highly reproducible. The Nextflow DSL2 implementation of this pipeline uses one container per process which makes it much easier to maintain and update software dependencies. Where possible, these processes have been submitted to and installed from nf-core/modules in order to make them available to all nf-core pipelines, and to everyone within the Nextflow community!
On release, automated continuous integration tests run the pipeline on a full-sized dataset on the AWS cloud infrastructure. This ensures that the pipeline runs on AWS, has sensible resource allocation defaults set to run on real-world datasets, and permits the persistent storage of results to benchmark between pipeline releases and other analysis sources. The results obtained from the full-sized test can be viewed on the nf-core website.
Pipeline summary
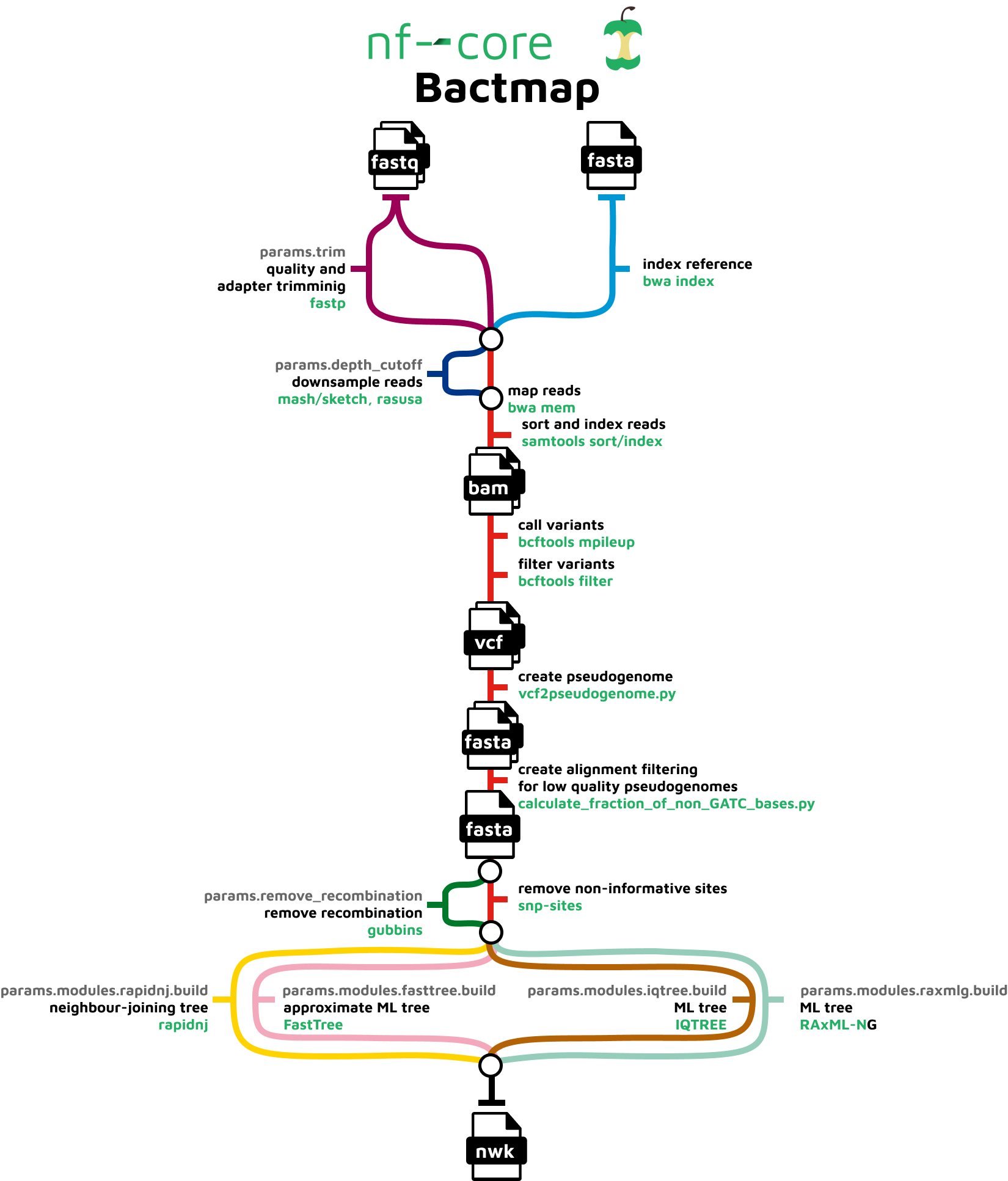
The pipeline is composed of the following steps:
- Index reference fasta file (
BWA index) - Trim reads for quality and adapter sequence (Optional) (
fastp) - Estimate genome size (
mash sketch) - Downsample fastq files (Optional) (
Rasusa) - Variant calling
- Read mapping (
BWA mem) - Sort and index alignments (
SAMtools) - Call and filter variants (
BCFtools) - Convert filtered bcf to pseudogenome fasta (
vcf2pseudogenome.py)
- Read mapping (
- Create alignment from pseudogenome by concatenating fasta files having first checked that the sample sequences are high quality(
calculate_fraction_of_non_GATC_bases.py) - Remove recombination (Optional) (
Gubbins) - Extract variant sites from alignment (
SNP-sites) - Construct phylogenetic tree (Optional)
Quick Start
-
Install
Nextflow(>=21.04.0) -
Install any of
Docker,Singularity,Podman,ShifterorCharliecloudfor full pipeline reproducibility (please only useCondaas a last resort; see docs) -
Download the pipeline and test it on a minimal dataset with a single command:
nextflow run nf-core/bactmap -profile test,<docker/singularity/podman/shifter/charliecloud/conda/institute>- Please check nf-core/configs to see if a custom config file to run nf-core pipelines already exists for your Institute. If so, you can simply use
-profile <institute>in your command. This will enable eitherdockerorsingularityand set the appropriate execution settings for your local compute environment. - If you are using
singularitythen the pipeline will auto-detect this and attempt to download the Singularity images directly as opposed to performing a conversion from Docker images. If you are persistently observing issues downloading Singularity images directly due to timeout or network issues then please use the--singularity_pull_docker_containerparameter to pull and convert the Docker image instead. Alternatively, it is highly recommended to use thenf-core downloadcommand to pre-download all of the required containers before running the pipeline and to set theNXF_SINGULARITY_CACHEDIRorsingularity.cacheDirNextflow options to be able to store and re-use the images from a central location for future pipeline runs. - If you are using
conda, it is highly recommended to use theNXF_CONDA_CACHEDIRorconda.cacheDirsettings to store the environments in a central location for future pipeline runs.
- Please check nf-core/configs to see if a custom config file to run nf-core pipelines already exists for your Institute. If so, you can simply use
-
Start running your own analysis!
nextflow run nf-core/bactmap -profile <docker/singularity/podman/conda/institute> --input samplesheet.csv --reference chromosome.fasta
Documentation
The nf-core/bactmap pipeline comes with documentation about the pipeline usage, parameters and output.
Credits
nf-core/bactmap was originally written by Anthony Underwood, Andries van Tonder and Thanh Le Viet.
We thank the following people for their extensive assistance in the development of this pipeline:
Anthony Underwood’s time working on the project was funded by the National Institute for Health Research
(NIHR) Global Health Research Unit for the Surveillance of Antimicrobial Resistance (Grant Reference Number 16/136/111)

Contributions and Support
If you would like to contribute to this pipeline, please see the contributing guidelines.
For further information or help, don’t hesitate to get in touch on the Slack #bactmap channel (you can join with this invite).
Citations
An extensive list of references for the tools used by the pipeline can be found in the CITATIONS.md file.
You can cite the nf-core publication as follows:
The nf-core framework for community-curated bioinformatics pipelines.
Philip Ewels, Alexander Peltzer, Sven Fillinger, Harshil Patel, Johannes Alneberg, Andreas Wilm, Maxime Ulysse Garcia, Paolo Di Tommaso & Sven Nahnsen.
Nat Biotechnol. 2020 Feb 13. doi: 10.1038/s41587-020-0439-x.