nf-core/metatdenovo
Assembly and annotation of metatranscriptomic or metagenomic data for prokaryotic, eukaryotic and viruses.
Introduction
nf-core/metatdenovo is a bioinformatics best-practice analysis pipeline for assembly and annotation of metatranscriptomic and metagenomic data from prokaryotes, eukaryotes or viruses.
On release, automated continuous integration tests run the pipeline on a full-sized dataset on the AWS cloud infrastructure. This ensures that the pipeline runs on AWS, has sensible resource allocation defaults set to run on real-world datasets, and permits the persistent storage of results to benchmark between pipeline releases and other analysis sources. The results obtained from the full-sized test can be viewed on the nf-core website.
Usage
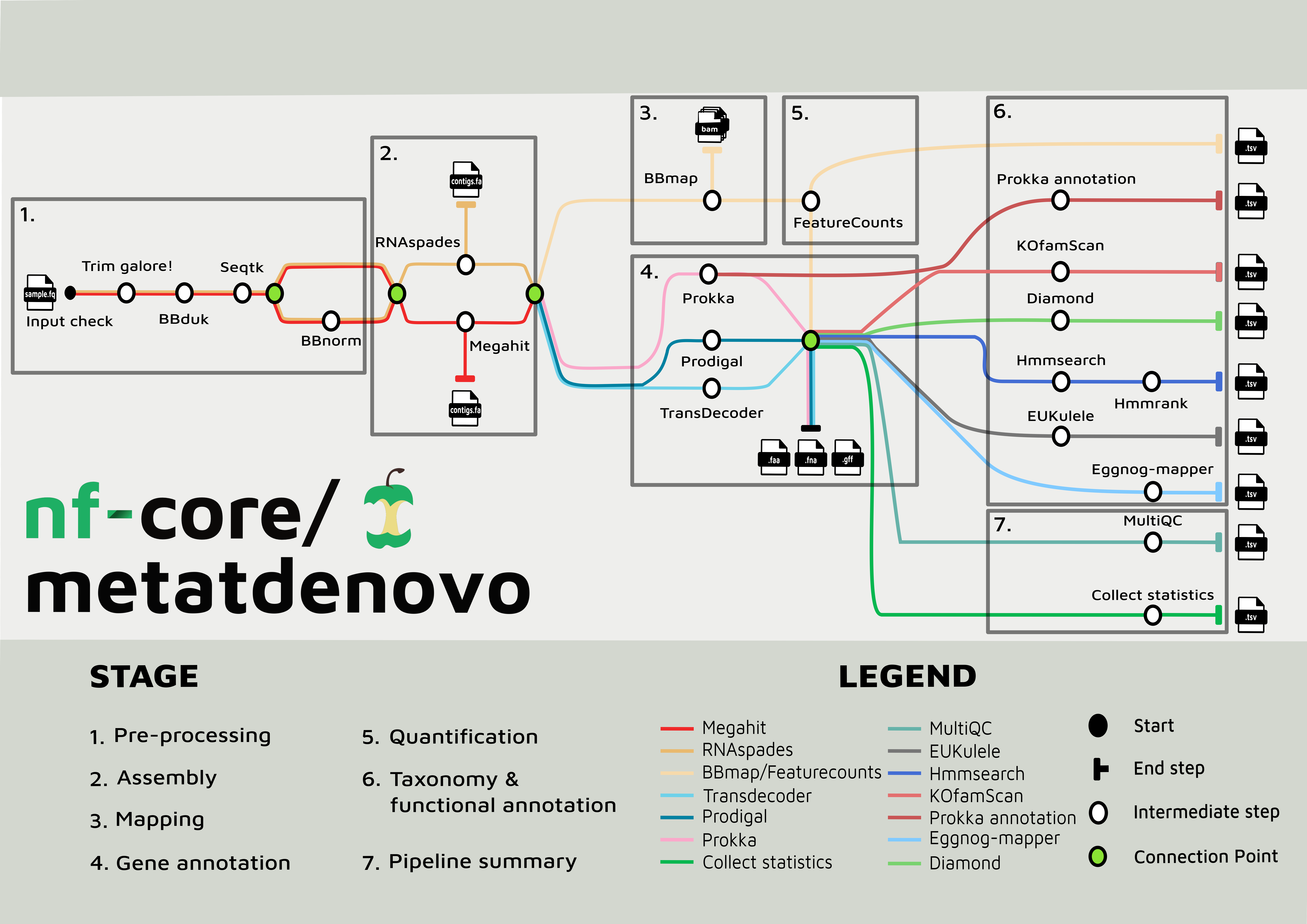
- Read QC (
FastQC) - Present QC for raw reads (
MultiQC) - Quality trimming and adapter removal for raw reads (
Trim Galore!) - Optional: Filter sequences with
BBduk - Optional: Normalize the sequencing depth with
BBnorm - Merge trimmed, pair-end reads (
Seqtk) - Choice of de novo assembly programs:
- Choice of orf caller:
TransDecodersuggested for eukaryotes; only ORFsProkkasuggested for prokaryotes; ORFs and other features plus functional annotationProdigalsuggested for Prokaryotes; only ORFs
- Quantification of genes identified in assemblies:
- Generate index of assembly (
BBmap index) - Mapping cleaned reads to the assembly for quantification (
BBmap) - Get raw counts per each gene present in the assembly (
Featurecounts) -> TSV table with collected featurecounts output
- Generate index of assembly (
- Functional annotation:
Prokkafeature identification and annotation for prokaryoteseggNOG-mapperKofamScanHMMERsearch ORFs with a set of HMM profiles, and rank results
- Taxonomic annotation:
- Summary statistics.
Usage
If you are new to Nextflow and nf-core, please refer to this page on how to set-up Nextflow. Make sure to test your setup with -profile test before running the workflow on actual data.
First, prepare a samplesheet with your input data that looks as follows:
samplesheet.csv:
sample,fastq_1,fastq_2
sample1,./data/S1_R1_001.fastq.gz,./data/S1_R2_001.fastq.gz
sample2,./data/S2_fw.fastq.gz,./data/S2_rv.fastq.gz
sample3,./S4x.fastq.gz,./S4y.fastq.gz
sample3,./a.fastq.gz,./b.fastq.gzEach row represents a fastq file (single-end) or a pair of fastq files (paired-end).
The fastq files need to end with .fq or .fastq, followed by .gz if gzipped.
Read files from multiple rows with the same sample name will be concatenated and treated as a single sample.
A mix of single-end and paired-end files is allowed, but do not mix single-end and paired-end for the same sample name.
Now, you can run the pipeline using:
nextflow run nf-core/metatdenovo \
-profile <docker/singularity/.../institute> \
--input samplesheet.csv \
--outdir <OUTDIR>Please provide pipeline parameters via the CLI or Nextflow -params-file option. Custom config files including those provided by the -c Nextflow option can be used to provide any configuration except for parameters; see docs.
For more details and further functionality, please refer to the usage documentation and the parameter documentation.
Pipeline output
To see the results of an example test run with a full size dataset refer to the results tab on the nf-core website pipeline page. For more details about the output files and reports, please refer to the output documentation.
Tables in the summary_tables directory under the output directory are made especially for further analysis in tools like R or Python.
Their formats are standardized and column names consistent between tables.
Credits
nf-core/metatdenovo was originally written by Danilo Di Leo (@danilodileo), Emelie Nilsson (@emnilsson) & Daniel Lundin (@erikrikarddaniel).
Contributions and Support
If you would like to contribute to this pipeline, please see the contributing guidelines.
For further information or help, don’t hesitate to get in touch on the Slack #metatdenovo channel (you can join with this invite).
Citations
If you use nf-core/metatdenovo for your analysis, please cite it using the following doi: 10.5281/zenodo.10666590
An extensive list of references for the tools used by the pipeline can be found in the CITATIONS.md file.
You can cite the nf-core publication as follows:
The nf-core framework for community-curated bioinformatics pipelines.
Philip Ewels, Alexander Peltzer, Sven Fillinger, Harshil Patel, Johannes Alneberg, Andreas Wilm, Maxime Ulysse Garcia, Paolo Di Tommaso & Sven Nahnsen.
Nat Biotechnol. 2020 Feb 13. doi: 10.1038/s41587-020-0439-x.