nf-core/nanostring
An analysis pipeline for Nanostring nCounter expression data.
Introduction
nf-core/nanostring is a bioinformatics pipeline that can be used to analyze NanoString data. The performed analysis steps include quality control and data normalization.
The pipeline is built using Nextflow, a workflow tool to run tasks across multiple compute infrastructures in a very portable manner. It uses Docker/Singularity containers making installation trivial and results highly reproducible. The Nextflow DSL2 implementation of this pipeline uses one container per process which makes it much easier to maintain and update software dependencies. Where possible, these processes have been submitted to and installed from nf-core/modules in order to make them available to all nf-core pipelines, and to everyone within the Nextflow community!
Pipeline summary
- Quality control with NACHO (
NACHO) - Perform normalization with NACHO
- Create count tables with provided metadata
- Present QC for NanoString data (
MultiQC)
Pipeline tubemap
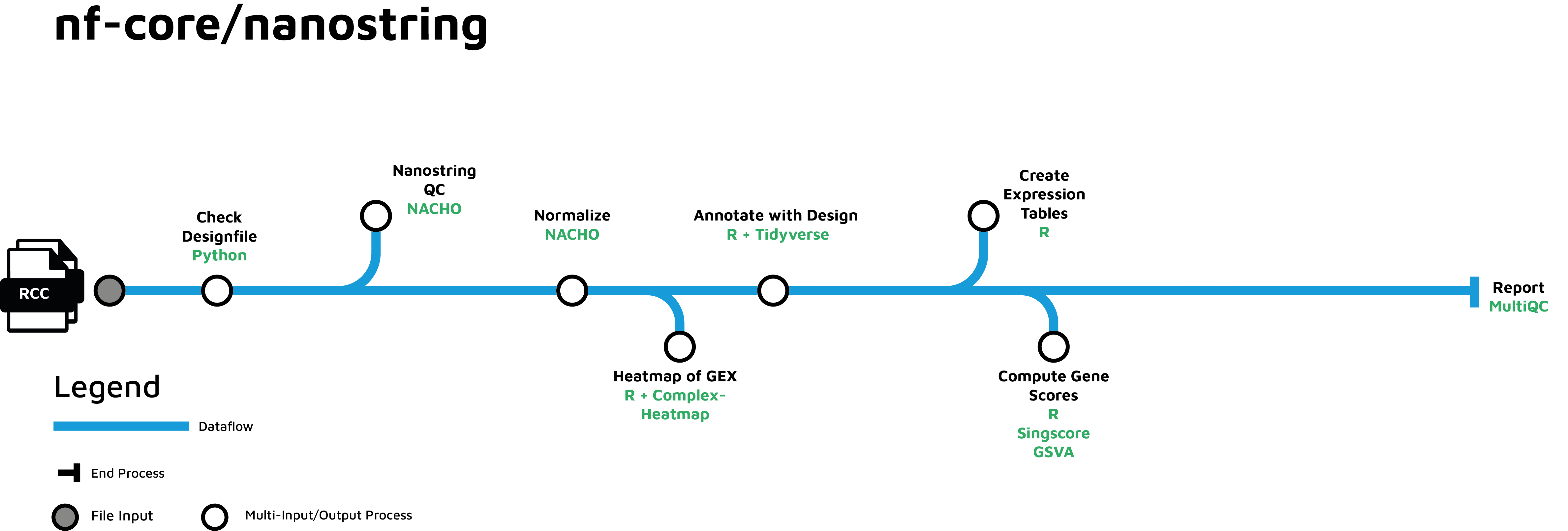
Usage
If you are new to Nextflow and nf-core, please refer to this page on how
to set-up Nextflow. Make sure to test your setup
with -profile test before running the workflow on actual data.
First, prepare a samplesheet with your input data that looks as follows:
samplesheet.csv:
RCC_FILE,RCC_FILE_NAME,SAMPLE_ID
/path/to/sample1.RCC,sample1.RCC,sample1
/path/to/sample2.RCC,sample2.RCC,sample2Each row represents a RCC file with counts.
Now, you can run the pipeline using:
nextflow run nf-core/nanostring \
-profile <docker/singularity/.../institute> \
--input samplesheet.csv \
--outdir <OUTDIR>Please provide pipeline parameters via the CLI or Nextflow -params-file option. Custom config files including those provided by the -c Nextflow option can be used to provide any configuration except for parameters; see docs.
For more details and further functionality, please refer to the usage documentation and the parameter documentation.
Pipeline output
To see the results of an example test run with a full size dataset refer to the results tab on the nf-core website pipeline page. For more details about the output files and reports, please refer to the output documentation.
Credits
nf-core/nanostring was originally written by Peltzer, Alexander & Mohr, Christopher. Extensive support was provided from other co-authors on the scientific or technical input required for the pipeline:
- Stadermann, Kai
- Zwick, Matthias
- Leparc, Germán
- Schmid, Ramona
Contributions and Support
If you would like to contribute to this pipeline, please see the contributing guidelines.
For further information or help, don’t hesitate to get in touch on the Slack #nanostring channel (you can join with this invite).
Citations
If you use nf-core/nanostring for your analysis, please cite the publication: nf-core/nanostring Bioinformatics and the Zenodo DOI.
An extensive list of references for the tools used by the pipeline can be found in the CITATIONS.md file.
You can cite the nf-core publication as follows:
The nf-core framework for community-curated bioinformatics pipelines.
Philip Ewels, Alexander Peltzer, Sven Fillinger, Harshil Patel, Johannes Alneberg, Andreas Wilm, Maxime Ulysse Garcia, Paolo Di Tommaso & Sven Nahnsen.
Nat Biotechnol. 2020 Feb 13. doi: 10.1038/s41587-020-0439-x.