nf-core/viralrecon
Assembly and intrahost/low-frequency variant calling for viral samples
Introduction
nf-core/viralrecon is a bioinformatics analysis pipeline used to perform assembly and intra-host/low-frequency variant calling for viral samples. The pipeline supports both Illumina and Nanopore sequencing data. For Illumina short-reads the pipeline is able to analyse metagenomics data typically obtained from shotgun sequencing (e.g. directly from clinical samples) and enrichment-based library preparation methods (e.g. amplicon-based: ARTIC SARS-CoV-2 enrichment protocol; or probe-capture-based). For Nanopore data the pipeline only supports amplicon-based analysis obtained from primer sets created and maintained by the ARTIC Network.
On release, automated continuous integration tests run the pipeline on a full-sized dataset on the AWS cloud infrastructure. This ensures that the pipeline runs on AWS, has sensible resource allocation defaults set to run on real-world datasets, and permits the persistent storage of results to benchmark between pipeline releases and other analysis sources. The results obtained from running the full-sized tests individually for each --platform option can be viewed on the nf-core website and the output directories will be named accordingly i.e. platform_illumina/ and platform_nanopore/.
The pipeline is built using Nextflow, a workflow tool to run tasks across multiple compute infrastructures in a very portable manner. It uses Docker/Singularity containers making installation trivial and results highly reproducible. The Nextflow DSL2 implementation of this pipeline uses one container per process which makes it much easier to maintain and update software dependencies. Where possible, these processes have been submitted to and installed from nf-core/modules in order to make them available to all nf-core pipelines, and to everyone within the Nextflow community!
Pipeline summary
The pipeline has numerous options to allow you to run only specific aspects of the workflow if you so wish. For example, for Illumina data you can skip the host read filtering step with Kraken 2 with --skip_kraken2 or you can skip all of the assembly steps with the --skip_assembly parameter. See the usage and parameter docs for all of the available options when running the pipeline.
The SRA download functionality has been removed from the pipeline (>=2.1) and ported to an independent workflow called nf-core/fetchngs. You can provide --nf_core_pipeline viralrecon when running nf-core/fetchngs to download and auto-create a samplesheet containing publicly available samples that can be accepted directly by the Illumina processing mode of nf-core/viralrecon.
A number of improvements were made to the pipeline recently, mainly with regard to the variant calling. Please see Major updates in v2.3 for a more detailed description.
Illumina
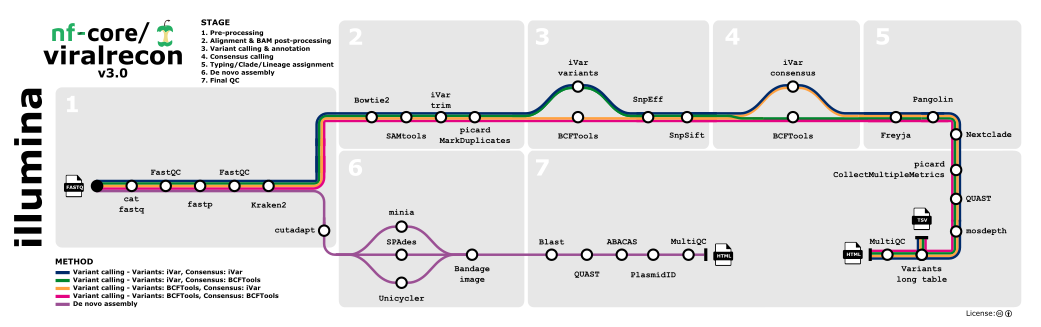
- Merge re-sequenced FastQ files (
cat) - Read QC (
FastQC) - Adapter trimming (
fastp) - Statistics/removal of host reads (
Kraken 2; optional) - Variant calling
- Read alignment (
Bowtie 2) - Sort and index alignments (
SAMtools) - Primer sequence removal (
iVar; amplicon data only) - Duplicate read marking (
picard; optional) - Alignment-level QC (
picard,SAMtools) - Genome-wide and amplicon coverage QC plots (
mosdepth) - Choice of multiple variant callers (
iVar variants; default for amplicon data ||BCFTools; default for metagenomics data) - Choice of multiple consensus callers (
BCFTools,BEDTools; default for both amplicon and metagenomics data ||iVar consensus) - Relative lineage abundance analysis from mixed SARS-CoV-2 samples (
Freyja) - Create variants long format table collating per-sample information for individual variants (
BCFTools), functional effect prediction (SnpSift) and lineage analysis (Pangolin)
- Read alignment (
- De novo assembly
- Present QC and visualisation for raw read, alignment, assembly and variant calling results (
MultiQC)
Nanopore
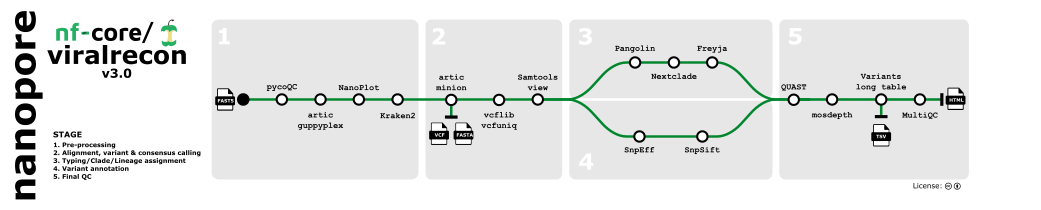
- Sequencing QC (
pycoQC) - Aggregate pre-demultiplexed reads from MinKNOW/Guppy (
artic guppyplex) - Read QC (
NanoPlot) - Statistics/removal of host reads (
Kraken 2; optional) - Align reads, call variants and generate consensus sequence (
artic minion) - Remove unmapped reads and obtain alignment metrics (
SAMtools) - Genome-wide and amplicon coverage QC plots (
mosdepth) - Downstream variant analysis:
- Count metrics (
BCFTools) - Variant annotation (
SnpEff,SnpSift) - Consensus assessment report (
QUAST) - Lineage analysis (
Pangolin) - Clade assignment, mutation calling and sequence quality checks (
Nextclade) - Recover relative lineage abundances from mixed SARS-CoV-2 samples (
Freyja) - Create variants long format table collating per-sample information for individual variants (
BCFTools), functional effect prediction (SnpSift) and lineage analysis (Pangolin)
- Count metrics (
- Present QC, visualisation and custom reporting for sequencing, raw reads, alignment and variant calling results (
MultiQC)
Usage
If you are new to Nextflow and nf-core, please refer to this page on how to set-up Nextflow. Make sure to test your setup with -profile test before running the workflow on actual data.
First, prepare a samplesheet with your input data that looks as follows:
samplesheet.csv:
sample,fastq_1,fastq_2
SAMPLE_1,AEG588A1_S1_L002_R1_001.fastq.gz,AEG588A1_S1_L002_R2_001.fastq.gzEach row represents a fastq file (single-end) or a pair of fastq files (paired end). Rows with the same sample identifier are considered technical replicates and merged automatically.
Typical commands
Illumina shotgun analysis
nextflow run nf-core/viralrecon \
--input samplesheet.csv \
--outdir <OUTDIR> \
--platform illumina \
--protocol metagenomic \
--genome 'MN908947.3' \
-profile -profile <docker/singularity/.../institute>Illumina amplicon analysis
nextflow run nf-core/viralrecon \
--input samplesheet.csv \
--outdir <OUTDIR> \
--platform illumina \
--protocol amplicon \
--genome 'MN908947.3' \
--primer_set artic \
--primer_set_version 3 \
--skip_assembly \
-profile -profile <docker/singularity/.../institute>Nanopore amplicon analysis:
nextflow run nf-core/viralrecon \
--input samplesheet.csv \
--outdir <OUTDIR> \
--platform nanopore \
--genome 'MN908947.3' \
--primer_set 'artic' \
--primer_set_version 3 \
--fastq_dir fastq_pass/ \
--sequencing_summary sequencing_summary.txt \
-profile -profile <docker/singularity/.../institute>Please provide pipeline parameters via the CLI or Nextflow -params-file option. Custom config files including those provided by the -c Nextflow option can be used to provide any configuration except for parameters; see docs.
For more details and further functionality, please refer to the usage documentation and the parameter documentation.
Automatic samplesheet generation
An executable Python script called fastq_dir_to_samplesheet.py has been provided if you are using --platform illumina and would like to auto-create an input samplesheet based on a directory containing FastQ files before you run the pipeline (requires Python 3 installed locally) e.g.
wget -L https://raw.githubusercontent.com/nf-core/viralrecon/master/bin/fastq_dir_to_samplesheet.py
./fastq_dir_to_samplesheet.py <FASTQ_DIR> samplesheet.csvReference genomes
You can find the default keys used to specify --genome in the genomes config file. This provides default options for
- Reference genomes (including SARS-CoV-2)
- Genome associates primer sets
- Nextclade datasets
The Pangolin and Nextclade lineage and clade definitions change regularly as new SARS-CoV-2 lineages are discovered. For instructions to use more recent versions of lineage analysis tools like Pangolin and Nextclade please refer to the updating containers section in the usage docs.
Where possible we are trying to collate links and settings for standard primer sets to make it easier to run the pipeline with standard keys; see usage docs.
Pipeline output
To see the results of an example test run with a full size dataset refer to the results tab on the nf-core website pipeline page. For more details about the output files and reports, please refer to the output documentation.
Credits
These scripts were originally written by Sarai Varona, Miguel Juliá, Erika Kvalem and Sara Monzon from BU-ISCIII and co-ordinated by Isabel Cuesta for the Institute of Health Carlos III, Spain. Through collaboration with the nf-core community the pipeline has now been updated substantially to include additional processing steps, to standardise inputs/outputs and to improve pipeline reporting; implemented and maintained primarily by Harshil Patel (@drpatelh) from Seqera Labs, Spain.
The key steps in the Nanopore implementation of the pipeline are carried out using the ARTIC Network’s field bioinformatics pipeline and were inspired by the amazing work carried out by contributors to the connor-lab/ncov2019-artic-nf pipeline originally written by Matt Bull for use by the COG-UK project. Thank you for all of your incredible efforts during this pandemic!
Many thanks to others who have helped out and contributed along the way too, including (but not limited to)*:
* Listed in alphabetical order
Contributions and Support
If you would like to contribute to this pipeline, please see the contributing guidelines.
For further information or help, don’t hesitate to get in touch on the Slack #viralrecon channel (you can join with this invite).
Citations
If you use nf-core/viralrecon for your analysis, please cite it using the following doi: 10.5281/zenodo.3901628
An extensive list of references for the tools used by the pipeline can be found in the CITATIONS.md file.
You can cite the nf-core publication as follows:
The nf-core framework for community-curated bioinformatics pipelines.
Philip Ewels, Alexander Peltzer, Sven Fillinger, Harshil Patel, Johannes Alneberg, Andreas Wilm, Maxime Ulysse Garcia, Paolo Di Tommaso & Sven Nahnsen.
Nat Biotechnol. 2020 Feb 13. doi: 10.1038/s41587-020-0439-x.