nf-core/bacass
Simple bacterial assembly and annotation pipeline
1.0.0). The latest
stable release is
2.6.0
.
Output
This document describes the output produced by the pipeline. Most of the plots are taken from the MultiQC report, which summarises results at the end of the pipeline.
Pipeline overview
The pipeline is built using Nextflow and processes data using the following steps:
- Quality trimming and QC of input reads
- Taxonomic classification of trimmed reads
- Assembly of trimmed reads
- Assembly visualization
- Assembly QC
- Annotation of the assembly
- Report describing results of (most of) the pipeline
Quality trimming and QC
This steps quality trims the end of reads, removes degenerate or too short reads and if needed, combines reads coming from multiple sequencing runs. It also runs FastQC which produces general quality metrics on your (trimmed) samples and plots them.
For further reading and documentation see the FastQC help.
Output directory: {sample}/{sample}_reads/
*.fastq.gz- trimmed (and combined reads)
*_fastqc.html- FastQC report, containing quality metrics for your trimmed reads
*_fastqc.zip- zip file containing the FastQC report, tab-delimited data file and plot images
FastQC screenshot
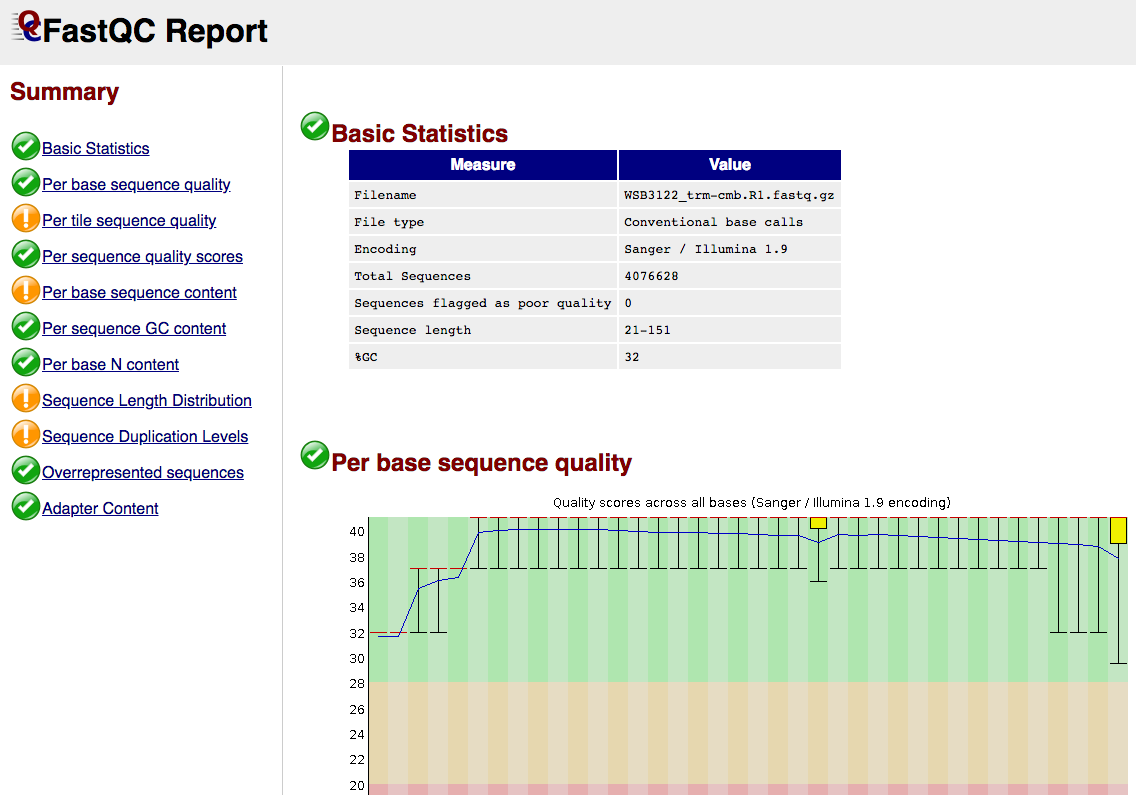
Taxonomic classification
This QC step classifies your reads using Kraken2 a k-mer based approach. This helps to identify samples that have purity issues. Ideally you will not want to assemble reads from samples that are contaminated or contain multiple species. If you like to visualize the report, try Pavian or Krakey.
Output directory: {sample}/
*_kraken2.report- Classification in the Kraken(1) report format. See webpage for more details
Kraken2 report screenshot
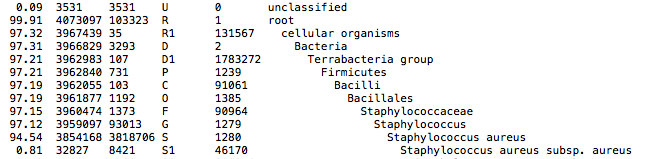
Assembly
Trimmed reads are assembled with Unicycler. Unicycler is a pipeline on its own, which at least for Illumina reads mainly acts as a frontend to Spades with added polishing steps.
Output directory: {sample}/
{sample}_assembly.fasta- Final assembly
{sample}_assembly.gfa- Final assembly in Graphical Fragment Assembly (GFA) format
{sample}_unicycler.log- Log file summarizing steps and intermediate results on the Unicycler execution
Assembly Visualization
The GFA file produced in the assembly step can be used to visualise the assembly graph, which is done here with Bandage. We highly recommend to run the Bandage GUI for more versatile visualisation options (annotations etc).
Output directory: {sample}/
{sample}_assembly.png- Bandage visualization of assembly
Bandage Screenshot
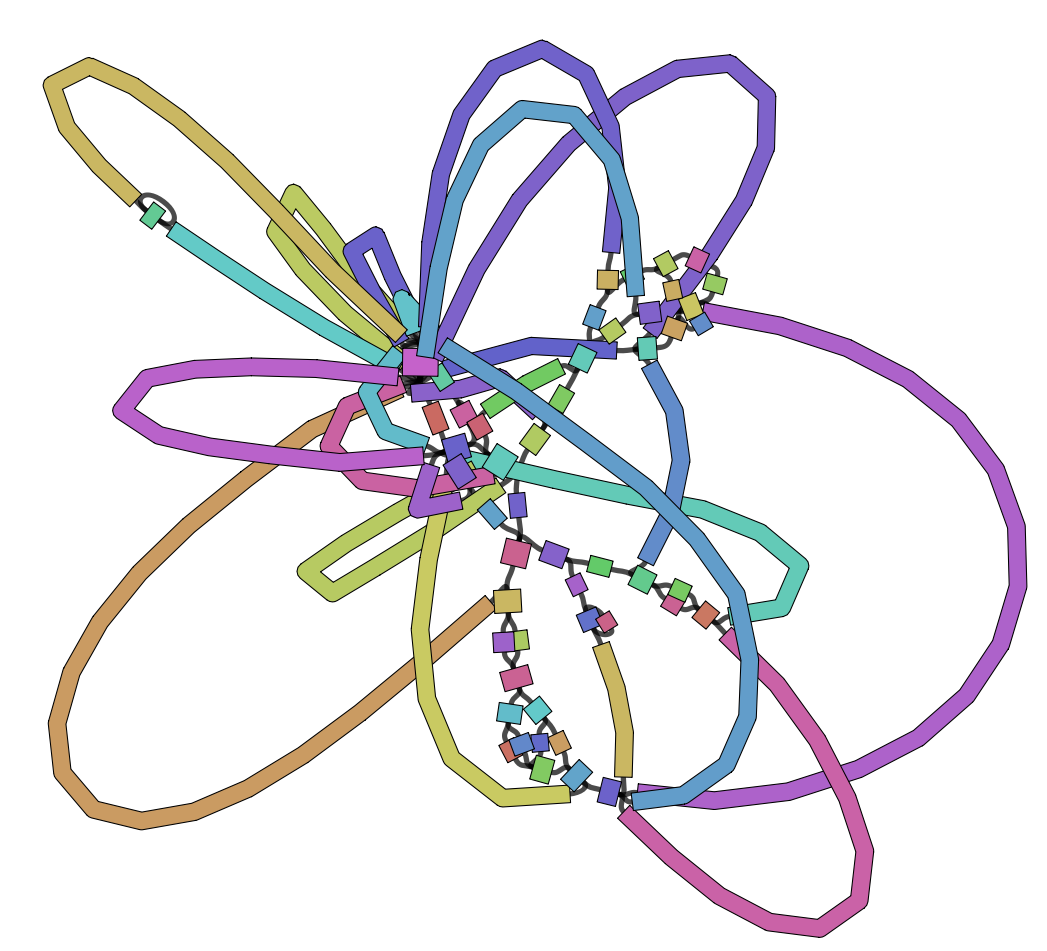

Assembly QC
The assembly QC is performed with QUAST. It reports multiple metrics including number of contigs, N50, lengths etc in form of an html report. It further creates an HTML file with integrated contig viewer (Icarus).
Output directory: {sample}/{sample}_assembly_QC
icarus.html- QUAST’s contig browser as HTML
report.html- QUAST assembly QC as HTML report
Quast Screenshot
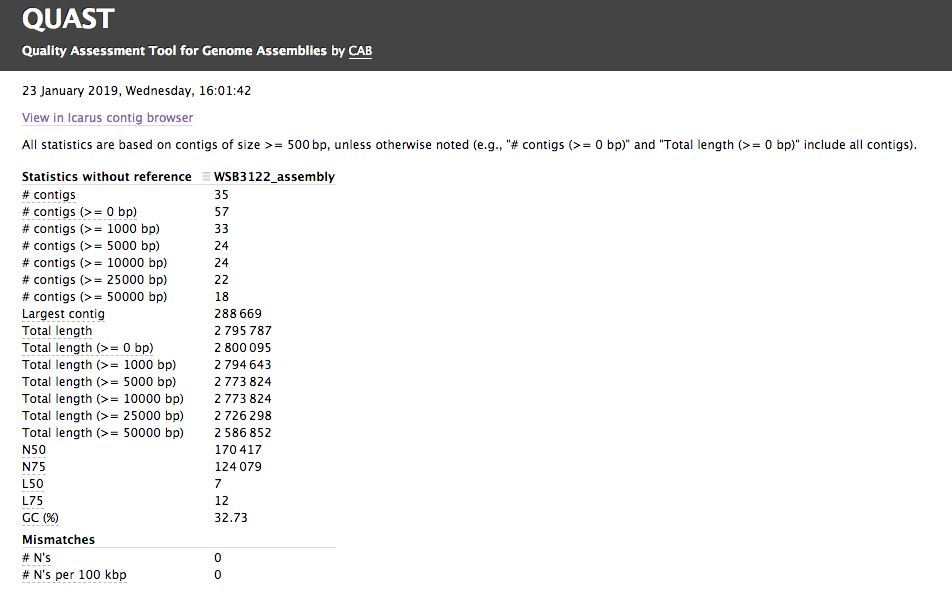
Icarus Screenshot
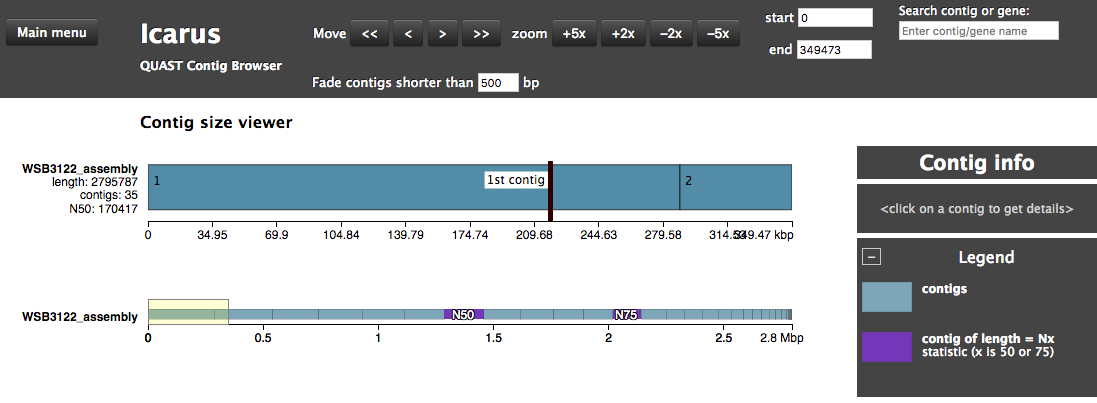
Annotation
The assembly is annotated with Prokka which acts as frontend for several annotation tools and includes rRNA and ORF predictions. See its documentation for a full description of all output files.
Output directory: {sample}/{sample}_annotation
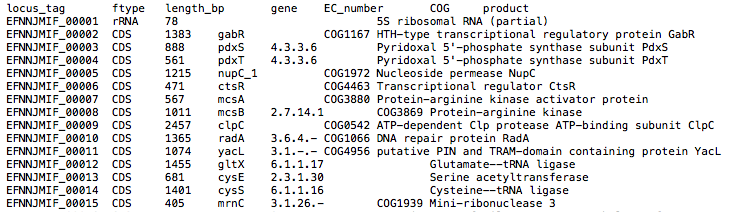
Report
Some pipeline results are visualised by MultiQC, which is a visualisation tool that generates a single HTML report summarising all samples in your project. Further statistics are available in within the report data directory.
The pipeline has special steps which allow the software versions used to be reported in the MultiQC output for future traceability.
Output directory: results/multiqc
Project_multiqc_report.html- MultiQC report - a standalone HTML file that can be viewed in your web browser
Project_multiqc_data/- Directory containing parsed statistics from the different tools used in the pipeline
For more information about how to use MultiQC reports, see http://multiqc.info