nf-core/molkart
A pipeline for processing Molecular Cartography data from Resolve Bioscience (combinatorial FISH)
Introduction
nf-core/molkart is a pipeline for processing Molecular Cartography data from Resolve Bioscience (combinatorial FISH). It takes as input a table of FISH spot positions (x,y,z,gene), a corresponding DAPI image (TIFF format) and optionally an additional staining image in the TIFF format. nf-core/molkart performs end-to-end processing of the data including image processing, QC filtering of spots, cell segmentation, spot-to-cell assignment and reports quality metrics such as the spot assignment rate, average spots per cell and segmentation mask size ranges.
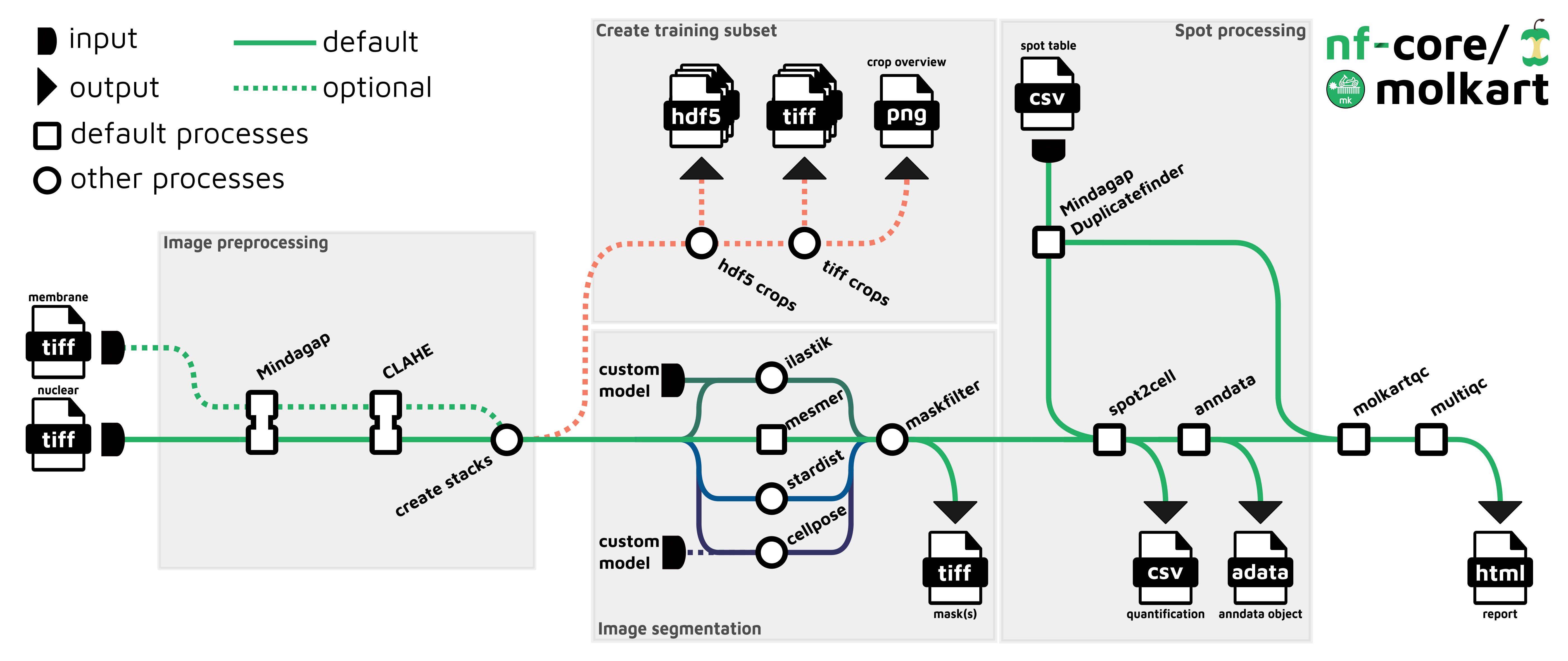
Image preprocessing
- Fill the grid pattern in provided images (
Mindagap) - Optionally apply contrast-limited adaptive histogram equalization
- If a second (membrane) image is present, combine images into a multichannel stack (if required for segmentation)
Cell segmentation
- Apply cell segmentation based on provided images, available options are: -
Cellpose-Mesmer-ilastik-Stardist - Filter cells based on cell size to remove artifacts
Spot processing
- Find duplicated spots near grid lines (
Mindagap) - Assign spots to segmented cells
Quality control
- Create quality-control metrics specific to this pipeline
- provide them to (
MultiQC) to create a report
Usage
If you are new to Nextflow and nf-core, please refer to this page on how
to set-up Nextflow. Make sure to test your setup
with -profile test before running the workflow on actual data.
First, prepare a samplesheet with your input data that looks as follows:
samplesheet.csv:
sample,nuclear_image,spot_locations,membrane_image
sample0,sample0_DAPI.tiff,sample0_spots.txt,sample0_WGA.tiffEach row represents an FOV (field-of-view). Columns represent the sample ID (all must be unique), the path to the respective nuclear image, the spot table, and optionally the path to the respective membrane image (or any additional image to improve segmentation).
Now, you can run the pipeline using all default values with:
nextflow run nf-core/molkart \
-profile <docker/singularity/.../institute> \
--input samplesheet.csv \
--outdir <OUTDIR>Please provide pipeline parameters via the CLI or Nextflow -params-file option. Custom config files including those provided by the -c Nextflow option can be used to provide any configuration except for parameters; see docs.
For more details and further functionality, please refer to the usage documentation and the parameter documentation.
Pipeline output
The pipeline outputs a matched cell-by-transcript table based on deduplicated spots and segmented cells, as well as preprocessing and segmentation intermediaries. To see the results of an example test run with a full size dataset refer to the results tab on the nf-core website pipeline page. For more details about the output files and reports, please refer to the output documentation.
Credits
nf-core/molkart was originally written by Kresimir Bestak, Florian Wuennemann.
We thank Maxime U Garcia for his assistance and support in the development of this pipeline.
Contributions and Support
If you would like to contribute to this pipeline, please see the contributing guidelines.
For further information or help, don’t hesitate to get in touch on the Slack #molkart channel (you can join with this invite).
Citations
If you use nf-core/molkart for your analysis, please cite it using the following doi: 10.5281/zenodo.10650749
An extensive list of references for the tools used by the pipeline can be found in the CITATIONS.md file.
You can cite the nf-core publication as follows:
The nf-core framework for community-curated bioinformatics pipelines.
Philip Ewels, Alexander Peltzer, Sven Fillinger, Harshil Patel, Johannes Alneberg, Andreas Wilm, Maxime Ulysse Garcia, Paolo Di Tommaso & Sven Nahnsen.
Nat Biotechnol. 2020 Feb 13. doi: 10.1038/s41587-020-0439-x.