nf-core/quantms
Quantitative mass spectrometry workflow. Currently supports proteomics experiments with complex experimental designs for DDA-LFQ, DDA-Isobaric and DIA-LFQ quantification.
This pipeline is archived and no longer maintained.
Archived pipelines can still be used, but may be outdated and will not receive bugfixes.
Introduction
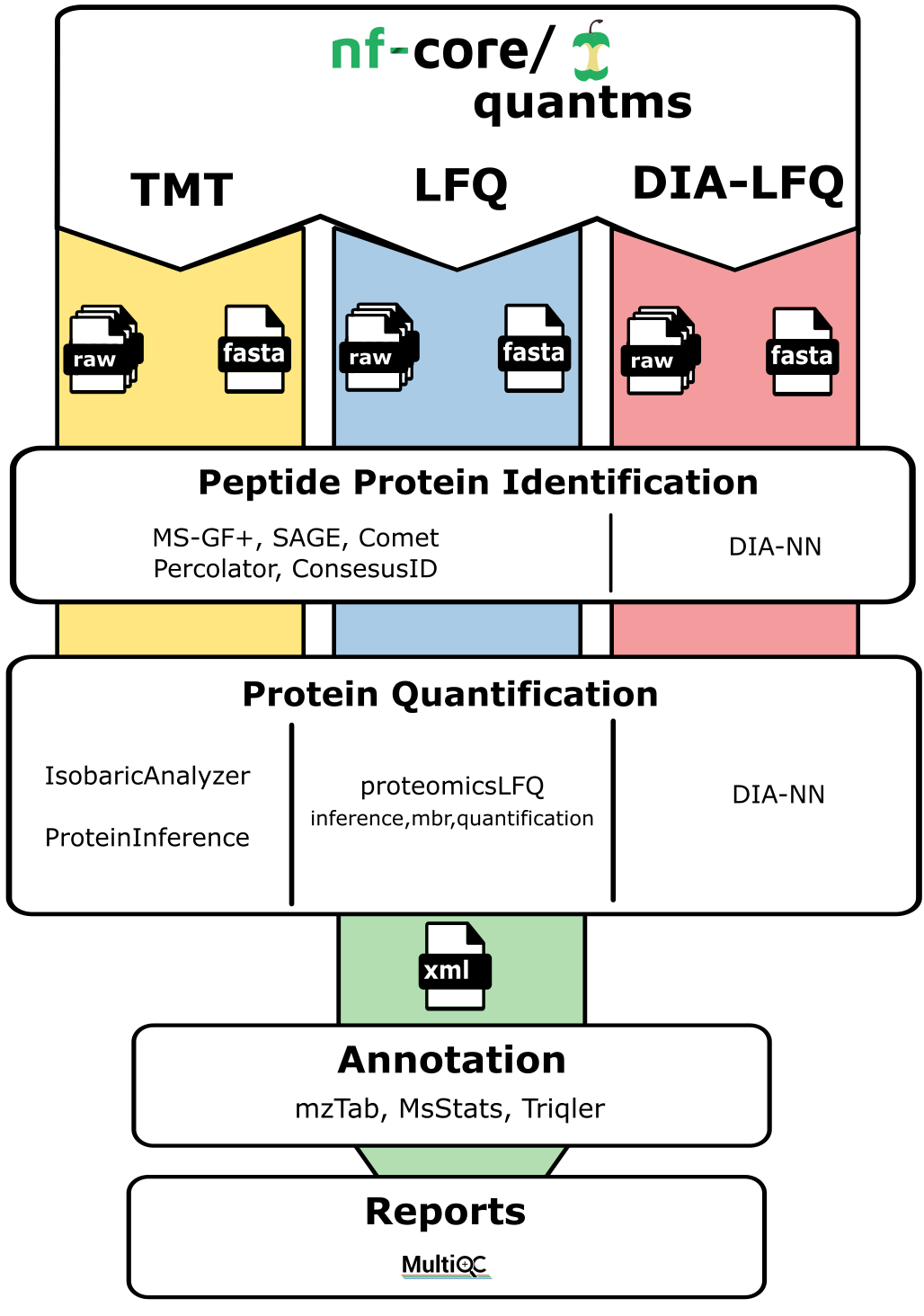
nf-core/quantms is a bioinformatics best-practice analysis pipeline for Quantitative Mass Spectrometry (MS). Currently, the workflow supports three major MS-based analytical methods: (i) Data dependant acquisition (DDA) label-free and Isobaric quantitation (e.g. TMT, iTRAQ); (ii) Data independent acquisition (DIA) label-free quantification (for details see our in-depth documentation on quantms).
The pipeline is built using Nextflow, a workflow tool to run tasks across multiple compute infrastructures in a very portable manner. It uses Docker/Singularity containers making installation trivial and results highly reproducible. The Nextflow DSL2 implementation of this pipeline uses one container per process which makes it much easier to maintain and update software dependencies. Where possible, these processes have been submitted to and installed from nf-core/modules in order to make them available to all nf-core pipelines, and to everyone within the Nextflow community!
The pipeline is built using Nextflow, a workflow tool to run tasks across multiple compute infrastructures in a very portable manner. It uses Docker/Singularity containers making installation trivial and results highly reproducible. The Nextflow DSL2 implementation of this pipeline uses one container per process which makes it much easier to maintain and update software dependencies. Where possible, these processes have been submitted to and installed from nf-core/modules in order to make them available to all nf-core pipelines, and to everyone within the Nextflow community!
On release, automated continuous integration tests run the pipeline on a full-sized dataset on the AWS cloud infrastructure. This ensures that the pipeline runs on AWS, has sensible resource allocation defaults set to run on real-world datasets, and permits the persistent storage of results to benchmark between pipeline releases and other analysis sources. The results obtained from the full-sized test can be viewed on the nf-core website. This gives you a hint on which reports and file types are produced by the pipeline in a standard run. The automatic continuous integration tests on every pull request evaluate different workflows, including peptide identification, quantification for LFQ, LFQ-DIA, and TMT test datasets.
Pipeline summary
nf-core/quantms allows uses to perform analyses of three main types of analytical mass spectrometry-based quantitative methods: DDA-LFQ, DDA-ISO, DIA-LFQ. Each of these workflows share some processes but also includes their own steps. In summary:
DDA-LFQ (data-dependent label-free quantification)
- RAW file conversion to mzML (
thermorawfileparser) - Peptide identification using
cometand/ormsgf+ - Re-scoring peptide identifications
percolator - Peptide identification FDR
openms fdr tool - Modification localization
luciphor - Quantification: Feature detection
proteomicsLFQ - Protein inference and quantification
proteomicsLFQ - QC report generation
pmultiqc - Normalization, imputation, significance testing with
MSstats
DDA-ISO (data-dependent quantification via isobaric labelling)
- RAW file conversion to mzML (
thermorawfileparser) - Peptide identification using
cometand/ormsgf+ - Re-scoring peptide identifications
percolator - Peptide identification FDR
openms fdr tool - Modification localization
luciphor - Extracts and normalizes isobaric labeling
IsobaricAnalyzer - Protein inference
ProteinInferenceorEpifanyfor bayesian inference. - Protein Quantification
ProteinQuantifier - QC report generation
pmultiqc - Normalization, imputation, significance testing with
MSstats
DIA-LFQ (data-independent label-free quantification)
- RAW file conversion to mzML when RAW as input(
thermorawfileparser) - Performing an optional step: Converting .d to mzML when bruker data as input and set
convert_dotdto true - DIA-NN analysis
dia-nn - Generation of output files (msstats)
- QC reports generation
pmultiqc
Functionality overview
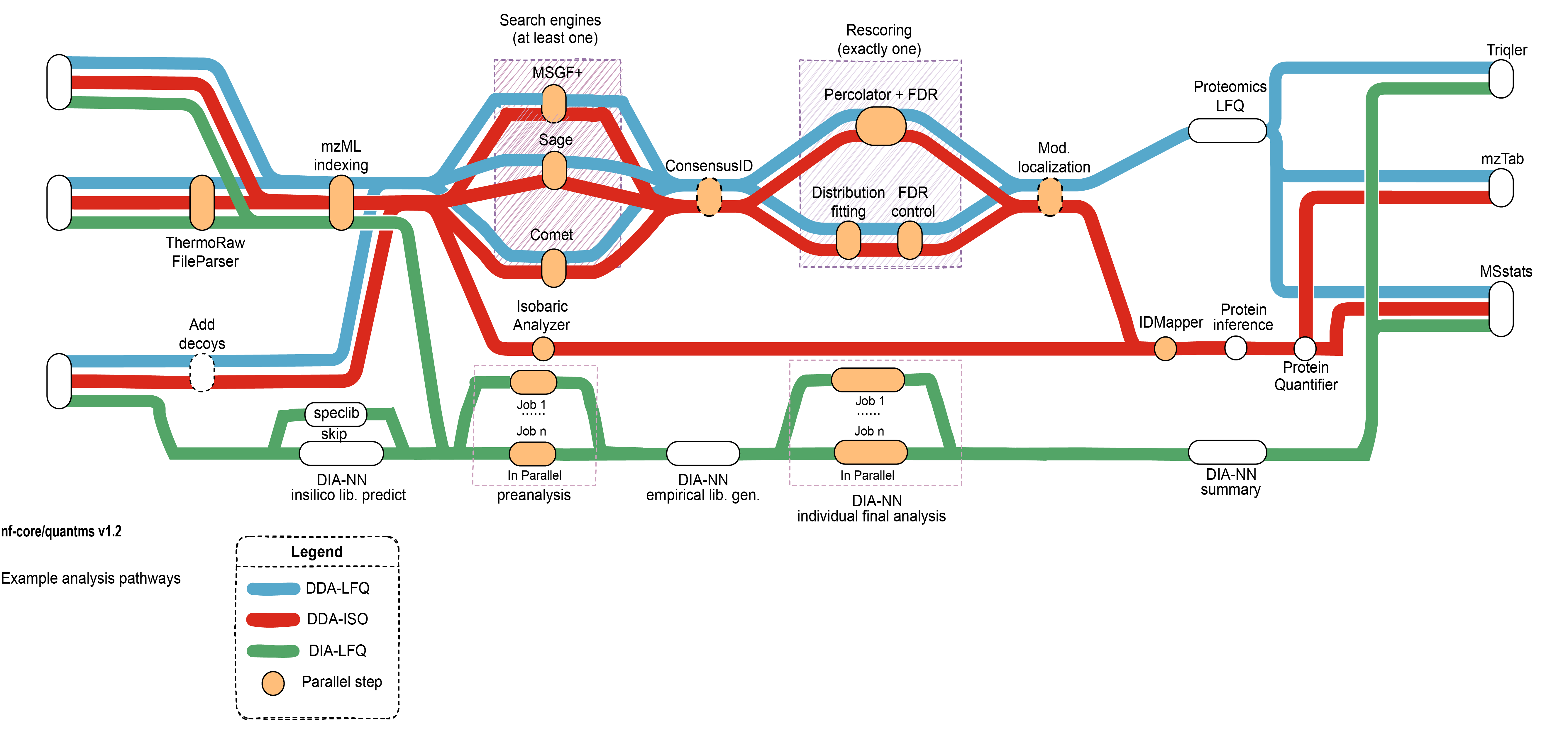
A graphical overview of suggested routes through the pipeline depending on context can be seen below.
Usage
If you are new to Nextflow and nf-core, please refer to this page on how to set-up Nextflow. Make sure to test your setup with -profile test before running the workflow on actual data.
First, find or create a sample-to-data relationship file (SDRF).
Have a look at public datasets that were already annotated here.
Those SDRFs should be ready for one-command re-analysis and you can just use the URL to the file on GitHub,
e.g., https://raw.githubusercontent.com/bigbio/proteomics-sample-metadata/master/annotated-projects/PXD000396/PXD000396.sdrf.tsv.
If you create your own, please adhere to the specifications and point the pipeline to your local folder or a remote location where you uploaded it to.
The second requirement is a protein sequence database. We suggest downloading a database for the organism(s)/proteins of interest from Uniprot.
Now, you can run the pipeline using:
nextflow run nf-core/quantms \
-profile <docker/singularity/.../institute> \
--input project.sdrf.tsv \
--database database.fasta \
--outdir <OUTDIR>Please provide pipeline parameters via the CLI or Nextflow -params-file option. Custom config files including those provided by the -c Nextflow option can be used to provide any configuration except for parameters;
see docs.
For more details and further functionality, please refer to the usage documentation and the parameter documentation.
Additional documentation and tutorial
The nf-core/quantms pipeline comes with a stand-alone full documentation including examples, benchmarks, and detailed explanation about the data analysis of proteomics data using quantms.
Pipeline output
To see the results of an example test run with a full size dataset refer to the results tab on the nf-core website pipeline page. For more details about the output files and reports, please refer to the output documentation.
Credits
nf-core/quantms was originally written by: Chengxin Dai (@daichengxin), Julianus Pfeuffer (@jpfeuffer) and Yasset Perez-Riverol (@ypriverol).
We thank the following people for their extensive assistance in the development of this pipeline:
- Timo Sachsenberg (@timosachsenberg)
- Wang Hong (@WangHong007)
Contributions and Support
If you would like to contribute to this pipeline, please see the contributing guidelines.
For further information or help, don’t hesitate to get in touch on the Slack #quantms channel (you can join with this invite).
Citations
An extensive list of references for the tools used by the pipeline can be found in the CITATIONS.md file.
You can cite the nf-core publication as follows:
The nf-core framework for community-curated bioinformatics pipelines.
Philip Ewels, Alexander Peltzer, Sven Fillinger, Harshil Patel, Johannes Alneberg, Andreas Wilm, Maxime Ulysse Garcia, Paolo Di Tommaso & Sven Nahnsen.
Nat Biotechnol. 2020 Feb 13. doi: 10.1038/s41587-020-0439-x.