nf-core/rnaseq
RNA sequencing analysis pipeline using STAR, RSEM, HISAT2 or Salmon with gene/isoform counts and extensive quality control.
3.13.2). The latest
stable release is
3.23.0
.
Introduction
nf-core/rnaseq is a bioinformatics pipeline that can be used to analyse RNA sequencing data obtained from organisms with a reference genome and annotation. It takes a samplesheet and FASTQ files as input, performs quality control (QC), trimming and (pseudo-)alignment, and produces a gene expression matrix and extensive QC report.
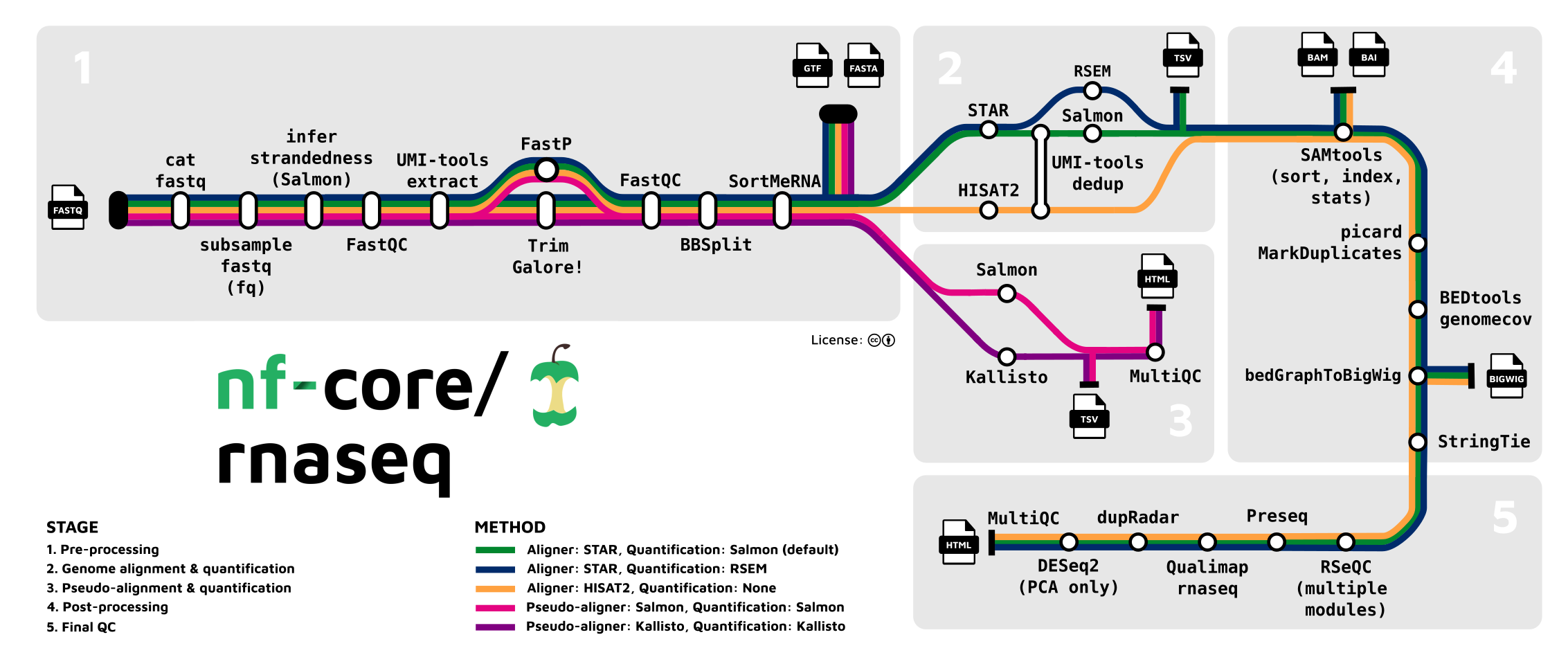
- Merge re-sequenced FastQ files (
cat) - Sub-sample FastQ files and auto-infer strandedness (
fq,Salmon) - Read QC (
FastQC) - UMI extraction (
UMI-tools) - Adapter and quality trimming (
Trim Galore!) - Removal of genome contaminants (
BBSplit) - Removal of ribosomal RNA (
SortMeRNA) - Choice of multiple alignment and quantification routes:
- Sort and index alignments (
SAMtools) - UMI-based deduplication (
UMI-tools) - Duplicate read marking (
picard MarkDuplicates) - Transcript assembly and quantification (
StringTie) - Create bigWig coverage files (
BEDTools,bedGraphToBigWig) - Extensive quality control:
- Pseudoalignment and quantification (
Salmonor ‘Kallisto’; optional) - Present QC for raw read, alignment, gene biotype, sample similarity, and strand-specificity checks (
MultiQC,R)
Note The SRA download functionality has been removed from the pipeline (
>=3.2) and ported to an independent workflow called nf-core/fetchngs. You can provide--nf_core_pipeline rnaseqwhen running nf-core/fetchngs to download and auto-create a samplesheet containing publicly available samples that can be accepted directly as input by this pipeline.
Warning Quantification isn’t performed if using
--aligner hisat2due to the lack of an appropriate option to calculate accurate expression estimates from HISAT2 derived genomic alignments. However, you can use this route if you have a preference for the alignment, QC and other types of downstream analysis compatible with the output of HISAT2.
Usage
If you are new to Nextflow and nf-core, please refer to this page on how
to set-up Nextflow. Make sure to test your setup
with -profile test before running the workflow on actual data.
First, prepare a samplesheet with your input data that looks as follows:
samplesheet.csv:
sample,fastq_1,fastq_2,strandedness
CONTROL_REP1,AEG588A1_S1_L002_R1_001.fastq.gz,AEG588A1_S1_L002_R2_001.fastq.gz,auto
CONTROL_REP1,AEG588A1_S1_L003_R1_001.fastq.gz,AEG588A1_S1_L003_R2_001.fastq.gz,auto
CONTROL_REP1,AEG588A1_S1_L004_R1_001.fastq.gz,AEG588A1_S1_L004_R2_001.fastq.gz,autoEach row represents a fastq file (single-end) or a pair of fastq files (paired end). Rows with the same sample identifier are considered technical replicates and merged automatically. The strandedness refers to the library preparation and will be automatically inferred if set to auto.
Warning: Please provide pipeline parameters via the CLI or Nextflow
-params-fileoption. Custom config files including those provided by the-cNextflow option can be used to provide any configuration except for parameters; see docs.
Now, you can run the pipeline using:
nextflow run nf-core/rnaseq \
--input samplesheet.csv \
--outdir <OUTDIR> \
--genome GRCh37 \
-profile <docker/singularity/.../institute>Please provide pipeline parameters via the CLI or Nextflow -params-file option. Custom config files including those
provided by the -c Nextflow option can be used to provide any configuration except for parameters;
see docs.
For more details and further functionality, please refer to the usage documentation and the parameter documentation.
Pipeline output
To see the results of an example test run with a full size dataset refer to the results tab on the nf-core website pipeline page. For more details about the output files and reports, please refer to the output documentation.
This pipeline quantifies RNA-sequenced reads relative to genes/transcripts in the genome and normalizes the resulting data. It does not compare the samples statistically in order to assign significance in the form of FDR or P-values. For downstream analyses, the output files from this pipeline can be analysed directly in statistical environments like R, Julia or via the nf-core/differentialabundance pipeline.
Online videos
A short talk about the history, current status and functionality on offer in this pipeline was given by Harshil Patel (@drpatelh) on 8th February 2022 as part of the nf-core/bytesize series.
You can find numerous talks on the nf-core events page from various topics including writing pipelines/modules in Nextflow DSL2, using nf-core tooling, running nf-core pipelines as well as more generic content like contributing to Github. Please check them out!
Credits
These scripts were originally written for use at the National Genomics Infrastructure, part of SciLifeLab in Stockholm, Sweden, by Phil Ewels (@ewels) and Rickard Hammarén (@Hammarn).
The pipeline was re-written in Nextflow DSL2 and is primarily maintained by Harshil Patel (@drpatelh) from Seqera Labs, Spain.
The pipeline workflow diagram was initially designed by Sarah Guinchard (@G-Sarah) and James Fellows Yates (@jfy133), further modifications where made by Harshil Patel (@drpatelh) and Maxime Garcia (@maxulysse).
Many thanks to other who have helped out along the way too, including (but not limited to):
- Alex Peltzer
- Colin Davenport
- Denis Moreno
- Edmund Miller
- Gregor Sturm
- Jacki Buros Novik
- Lorena Pantano
- Matthias Zepper
- Maxime Garcia
- Olga Botvinnik
- @orzechoj
- Paolo Di Tommaso
- Rob Syme
Contributions and Support
If you would like to contribute to this pipeline, please see the contributing guidelines.
For further information or help, don’t hesitate to get in touch on the Slack #rnaseq channel (you can join with this invite).
Citations
If you use nf-core/rnaseq for your analysis, please cite it using the following doi: 10.5281/zenodo.1400710
An extensive list of references for the tools used by the pipeline can be found in the CITATIONS.md file.
You can cite the nf-core publication as follows:
The nf-core framework for community-curated bioinformatics pipelines.
Philip Ewels, Alexander Peltzer, Sven Fillinger, Harshil Patel, Johannes Alneberg, Andreas Wilm, Maxime Ulysse Garcia, Paolo Di Tommaso & Sven Nahnsen.
Nat Biotechnol. 2020 Feb 13. doi: 10.1038/s41587-020-0439-x.