nf-core/sarek
Analysis pipeline to detect germline or somatic variants (pre-processing, variant calling and annotation) from WGS / targeted sequencing
2.5.2). The latest
stable release is
3.8.1
.
Introduction
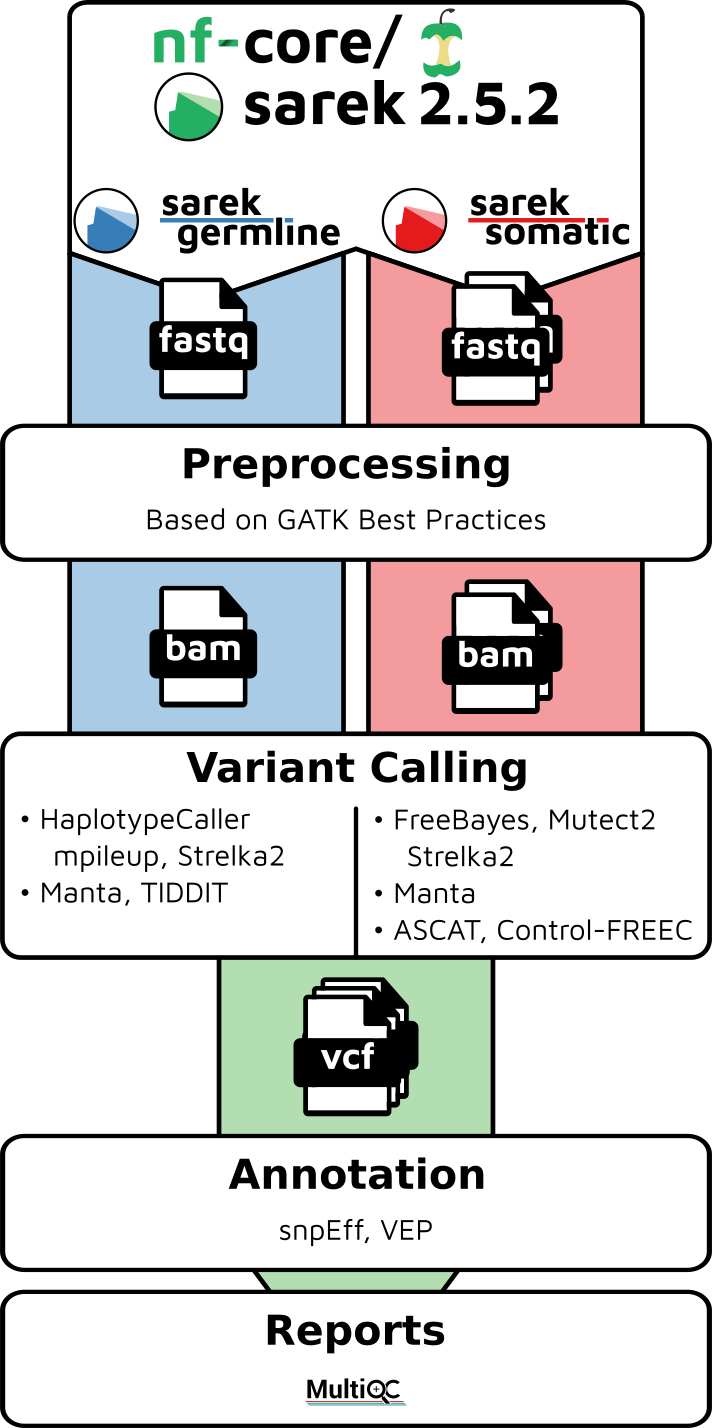
Sarek is a workflow designed to run analyses on whole genome or targeted sequencing data from regular samples or tumour / normal pairs and could include additional relapses.
It’s built using Nextflow, a domain specific language for workflow building, across multiple compute infrastructures in a very portable manner. Software dependencies are handled using Conda, Docker or Singularity - environment/container technologies that provide excellent reproducibility and ease of use. Thus making installation trivial and results highly reproducible.
It’s listed on the Elixir - Tools and Data Services Registry, Dockstore and omicX - Bioinformatics tools.
Documentation
The nf-core/sarek pipeline comes with documentation about the pipeline, found in the docs/ directory:
- Installation
- Pipeline configuration
- Running the pipeline
- Output and how to interpret the results
- Troubleshooting
Credits
Sarek was developed at the National Genomics Infastructure and National Bioinformatics Infastructure Sweden which are both platforms at SciLifeLab, with the support of The Swedish Childhood Tumor Biobank (Barntumörbanken).
Main authors:
Helpful contributors:
- Johannes Alneberg
- Phil Ewels
- Jesper Eisfeldt
- Malin Larsson
- Marcel Martin
- Alexander Peltzer
- Nilesh Tawari
- arontommi
- bjornnystedt
- gulfshores
- KochTobi
- pallolason
- Sebastian-D
- silviamorins
Contributions & Support
If you would like to contribute to this pipeline, please see the contributing guidelines.
For further information or help, don’t hesitate to get in touch on Slack (you can join with this invite) or contact us: maxime.garcia@scilifelab.se, szilveszter.juhos@scilifelab.se
CHANGELOG
Aknowledgements
Citation
If you use nf-core/sarek for your analysis, please cite the Sarek pre-print as follows:
Garcia MU, Juhos S, Larsson M, Olason PI, Martin M, Eisfeldt J, DiLorenzo S, Sandgren J, de Ståhl TD, Wirta V, Nistér M, Nystedt B, Käller M. Sarek: A portable workflow for whole-genome sequencing analysis of germline and somatic variants. bioRxiv. 2018. p. 316976. doi: 10.1101/316976.
You can cite the sarek zenodo record for a specific version using the following doi: 10.5281/zenodo.3476426
You can cite the nf-core pre-print as follows:
Ewels PA, Peltzer A, Fillinger S, Alneberg JA, Patel H, Wilm A, Garcia MU, Di Tommaso P, Nahnsen S. nf-core: Community curated bioinformatics pipelines. bioRxiv. 2019. p. 610741. doi: 10.1101/610741.